Measurements overview
Some tips & examples for doing measurements in #molstar at https://molstar.org/viewer
- label, distance, angle, dihedral
- orientation (box/axes from principal components)
- plane (best fit)
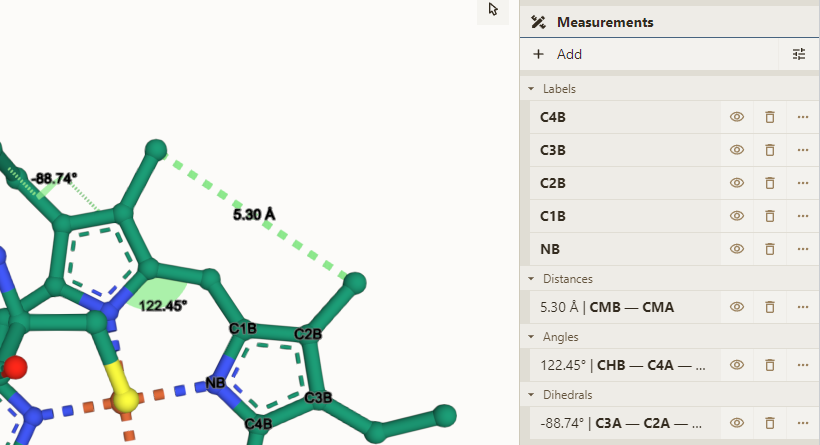
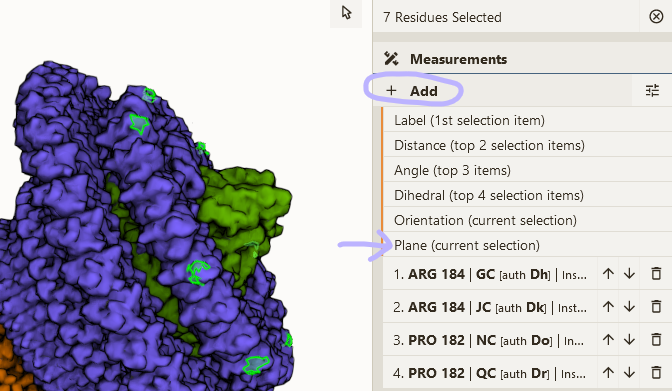
Basic measurements
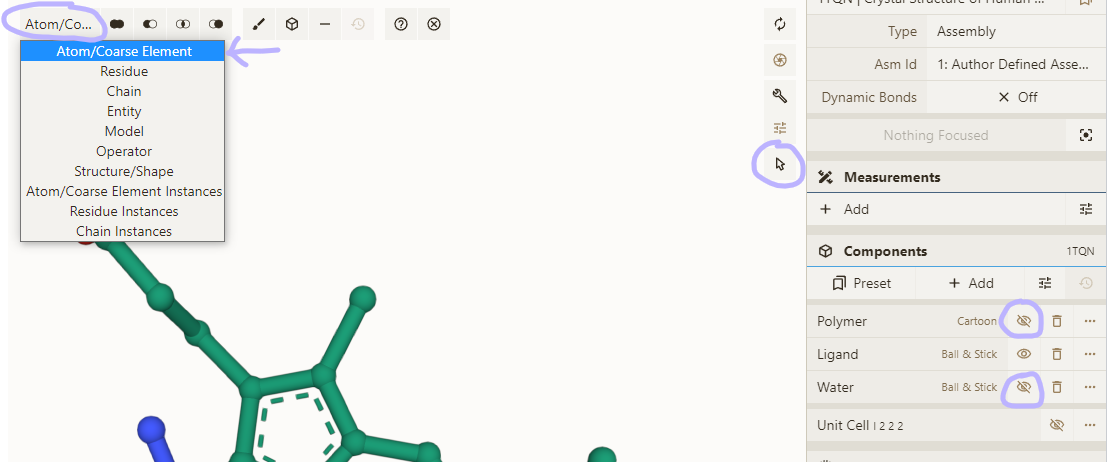
- Load PDB 1TQN; hide Polymer and Water component
- Enable selection mode; set granularity to "Atom/Element"
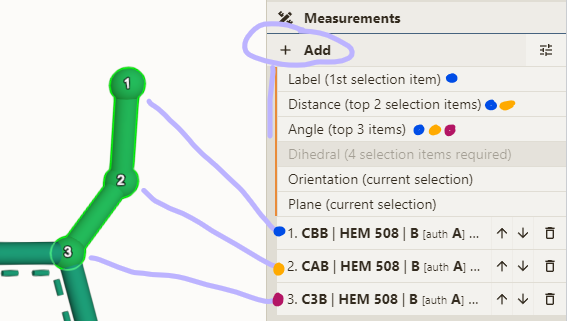
- Open Measurements panel & select some atoms
- Click one of the options available (depend on number of selected atoms)
View: https://molstar.org/viewer/?snapshot-url=https://molstar.org/viewer-docs/tips/measurements/basic.molx&snapshot-url-type=molx
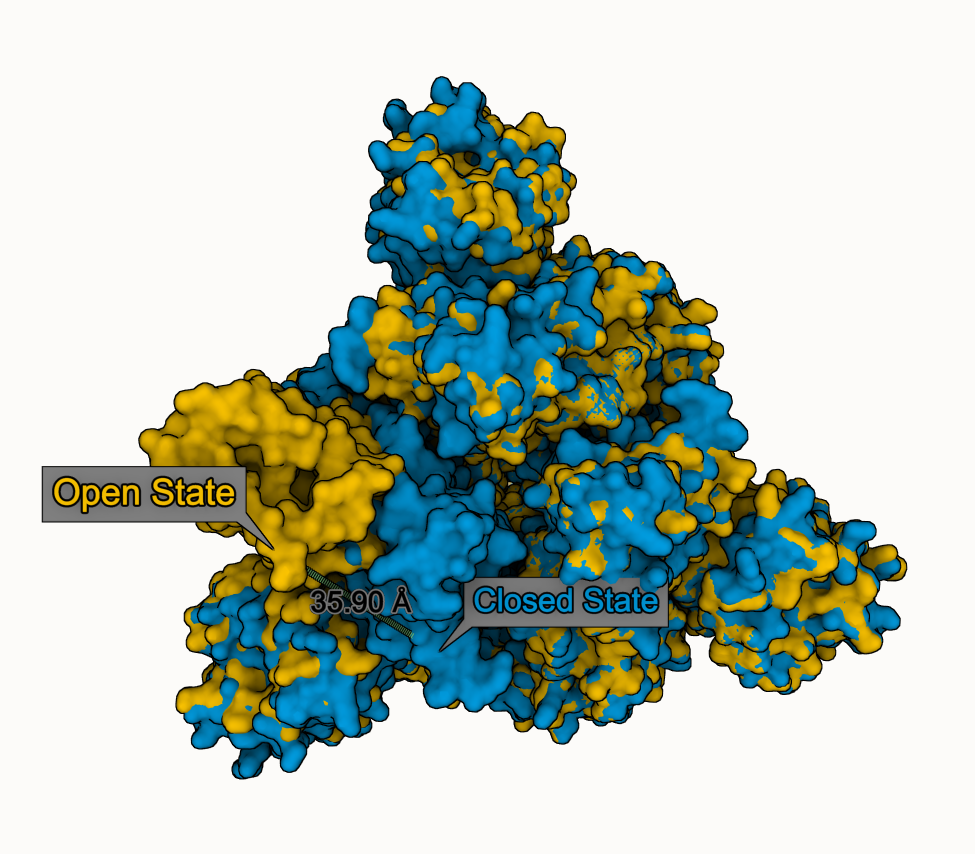
Distance example
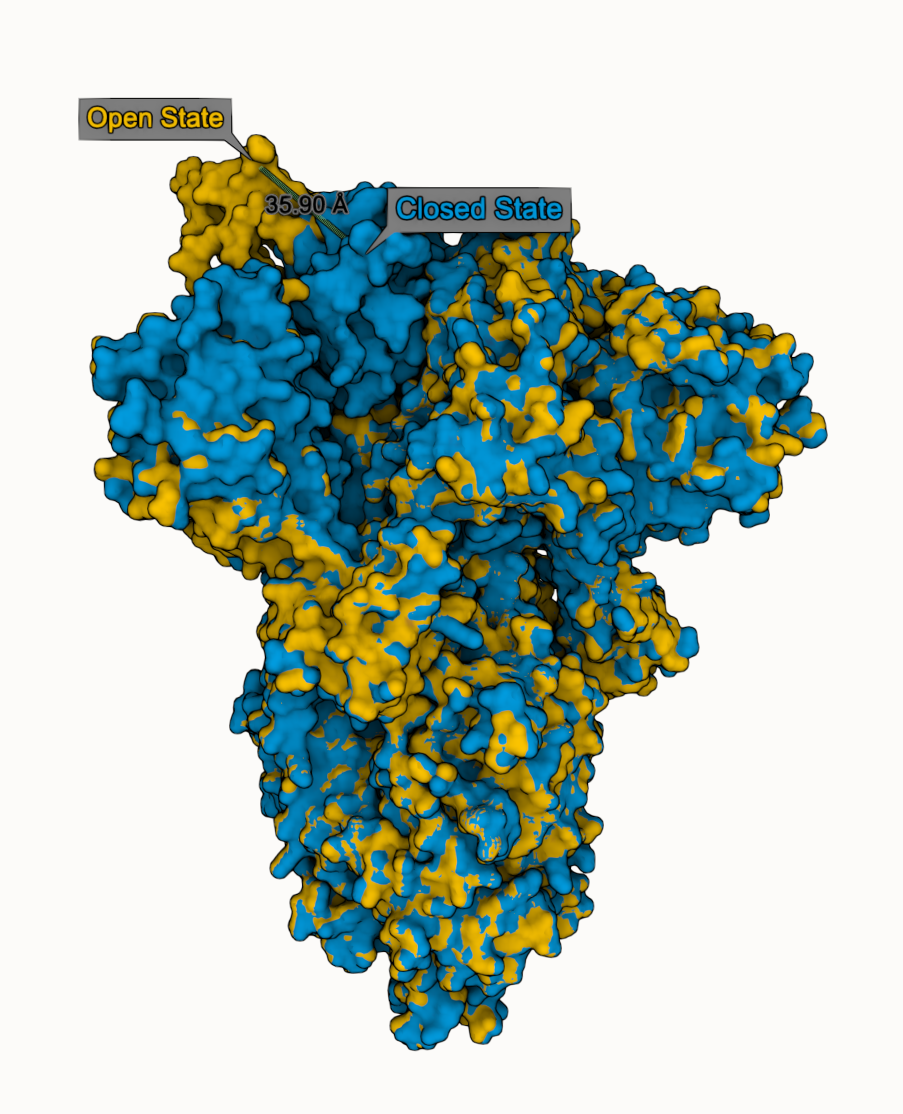
We compare closed (PDB 6VXX) and open (PDB 6VYB) state of SARS-CoV-2 spike glycoprotein
- Superpose both states
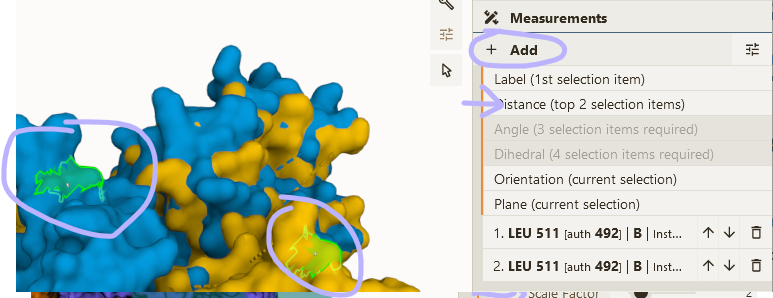
- Select Leu 511 from chain B in both states
- Add "distance" & "label" using Measurements panel
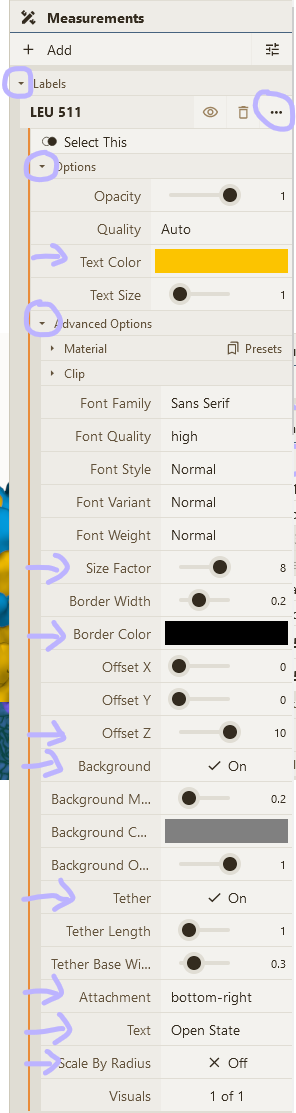
- Adjust visual options
View: https://molstar.org/viewer/?snapshot-url=https://molstar.org/viewer-docs/tips/measurements/distance.molx&snapshot-url-type=molx

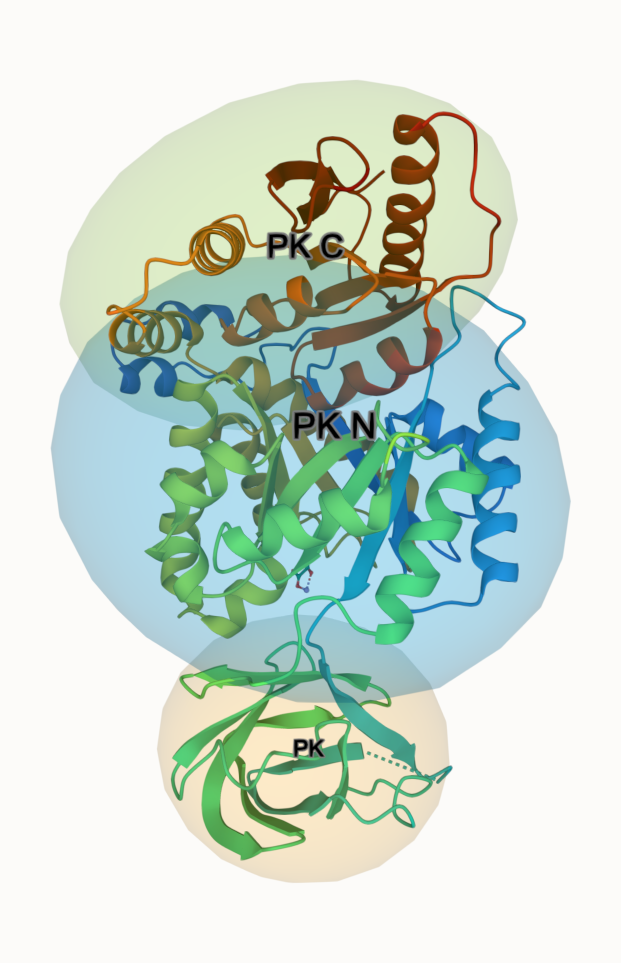
Orientation box/axes
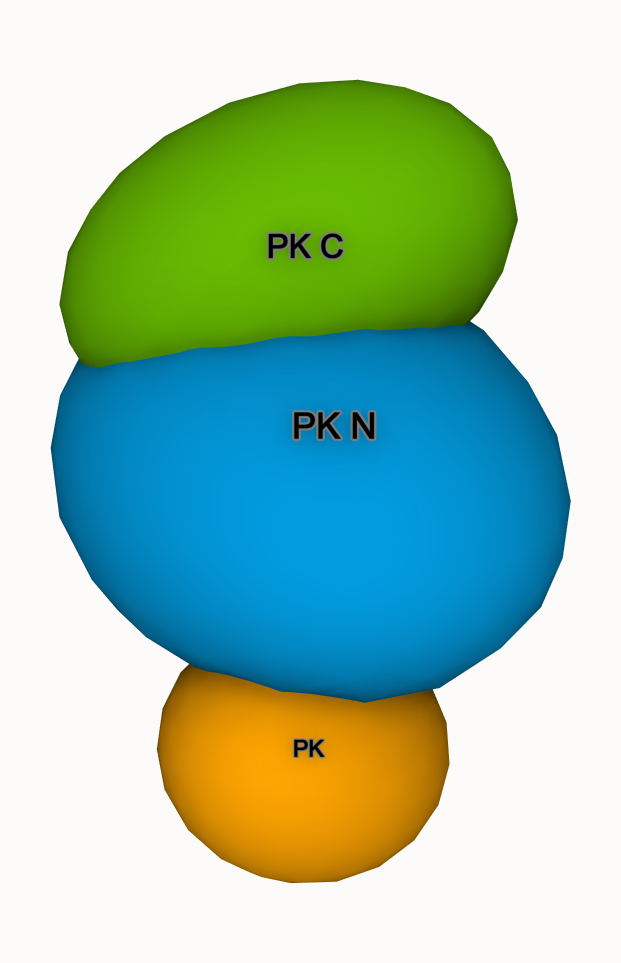
We use "orientation" ellipsoids to show the 3 domains of Pyruvate Kinase (PDB 1PKN)
- Select the PK domain (116-217)
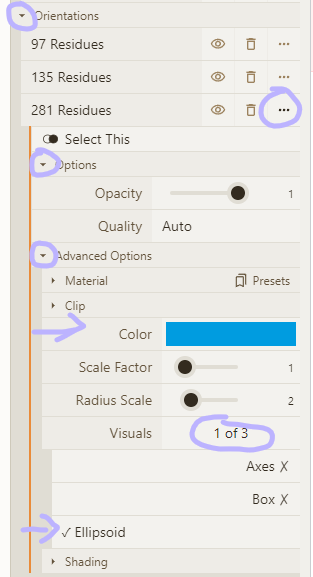
- Add "orientation" and "label"; adjust visual options
- Repeat for PK N (12-115 & 218-395) and PK C (396-530)
View: https://molstar.org/viewer/?snapshot-url=https://molstar.org/viewer-docs/tips/measurements/orientation.molx&snapshot-url-type=molx
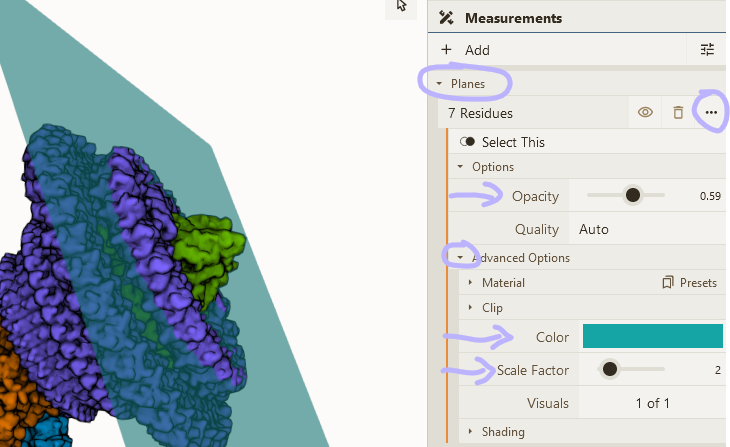
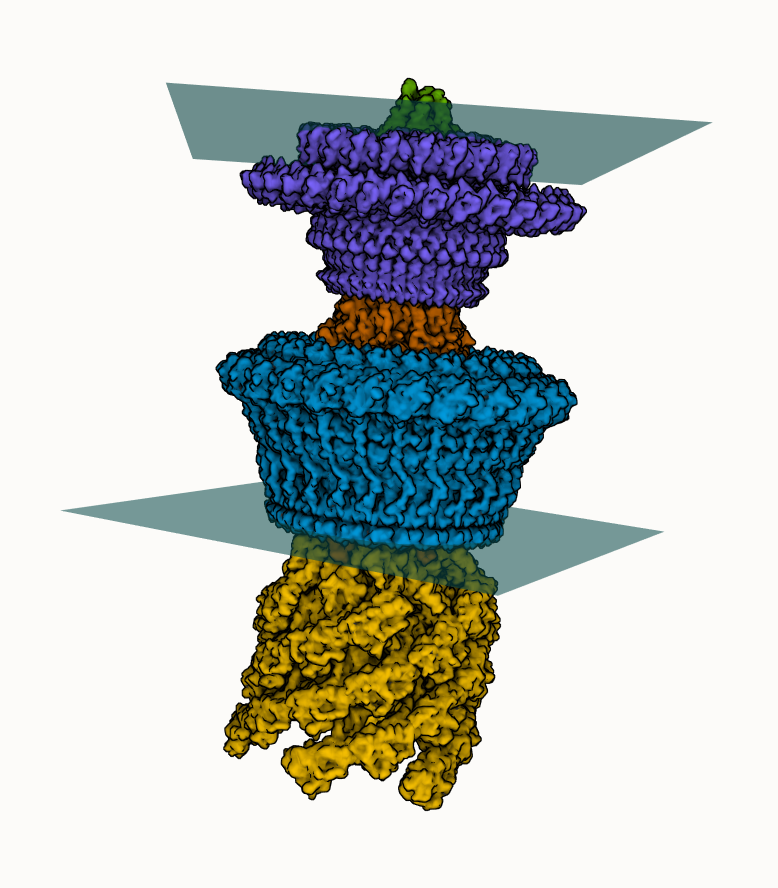
Best fit plane
We can add two best fit planes indicating lipid membranes to the bacterial flagellar motor-hook complex (@Jiaxing_Tan_ et al.)
- Select a few residues at the edge of the M-ring protein
- Add "plane" using Measurements panel
- Adjust visual options
- Repeat for the L-ring protein
View: https://molstar.org/viewer/?snapshot-url=https://molstar.org/viewer-docs/tips/measurements/plane.molx&snapshot-url-type=molx